Written by ITW
September 29, 2021
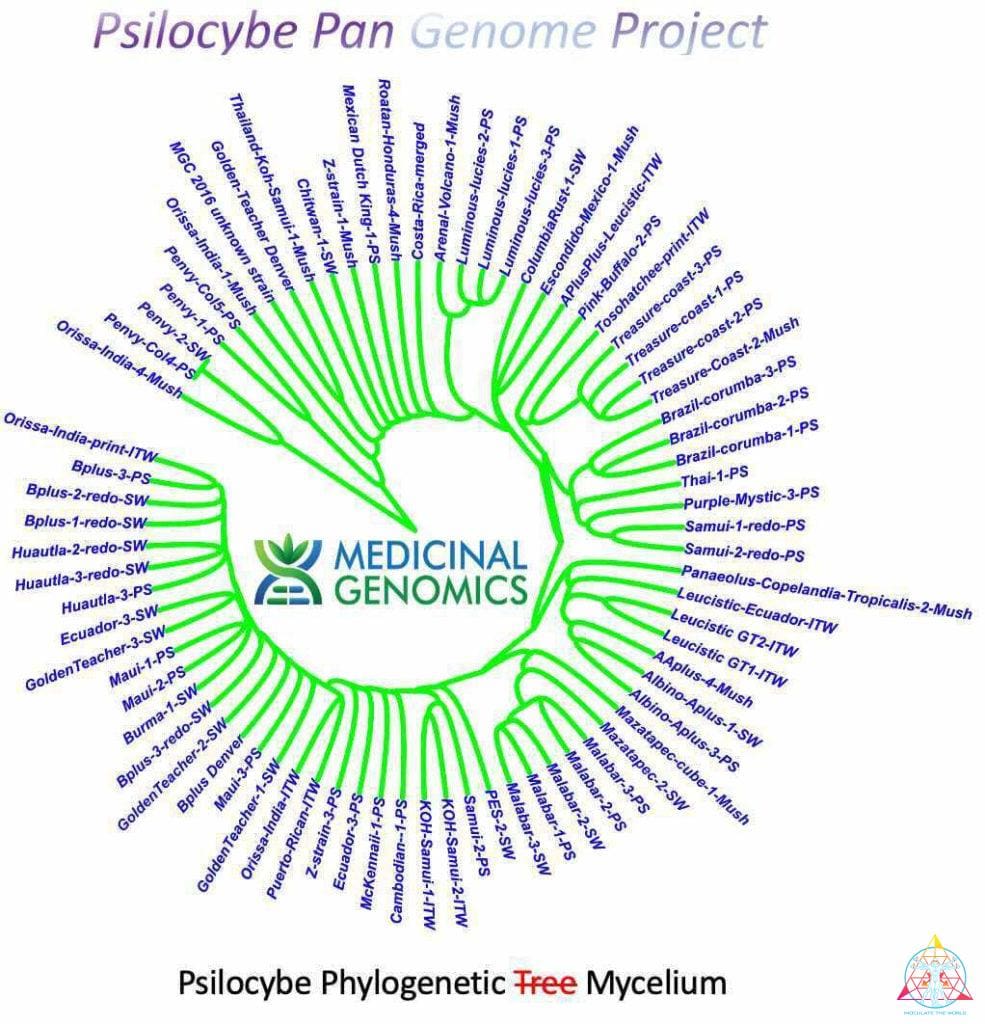
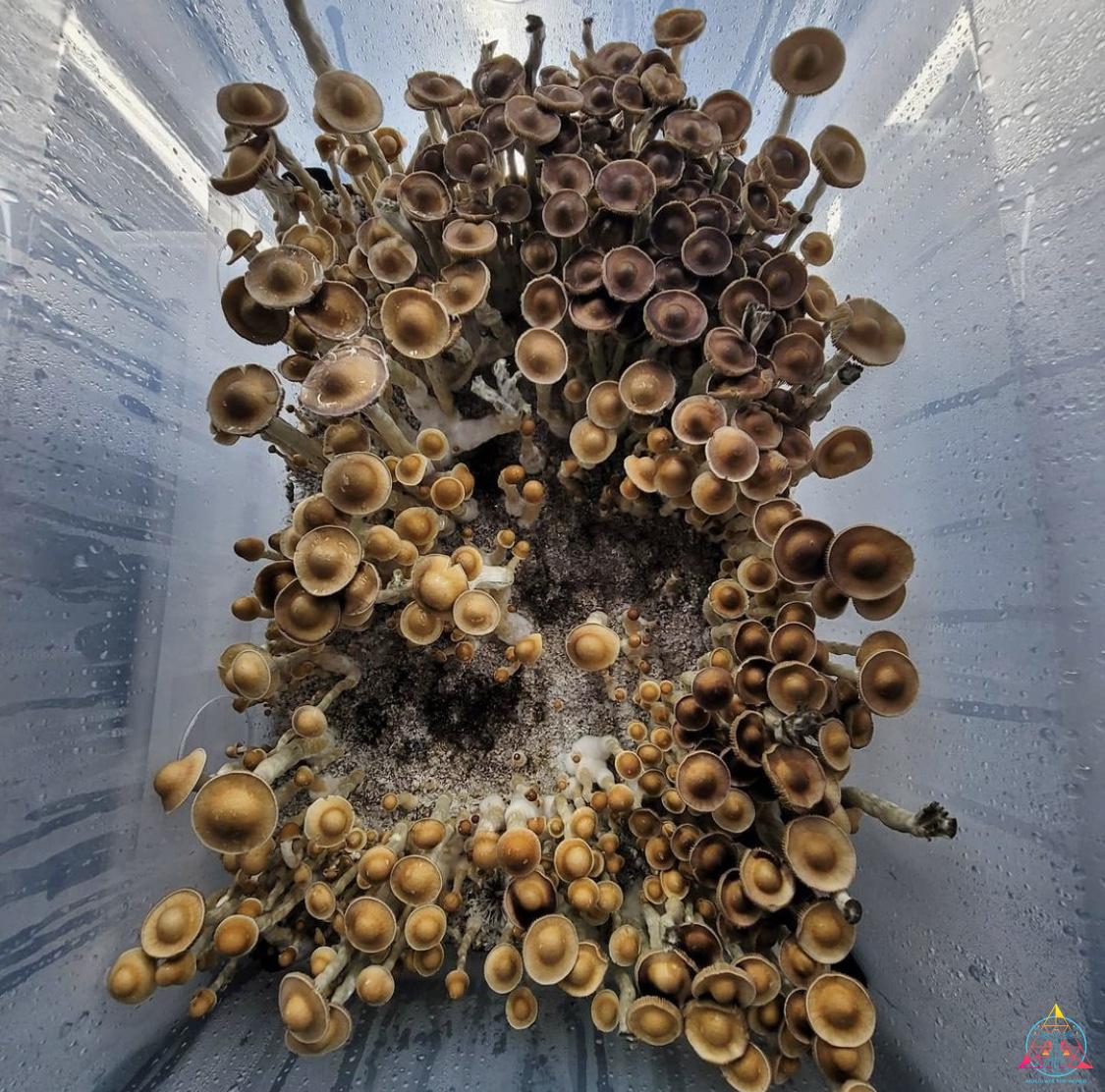
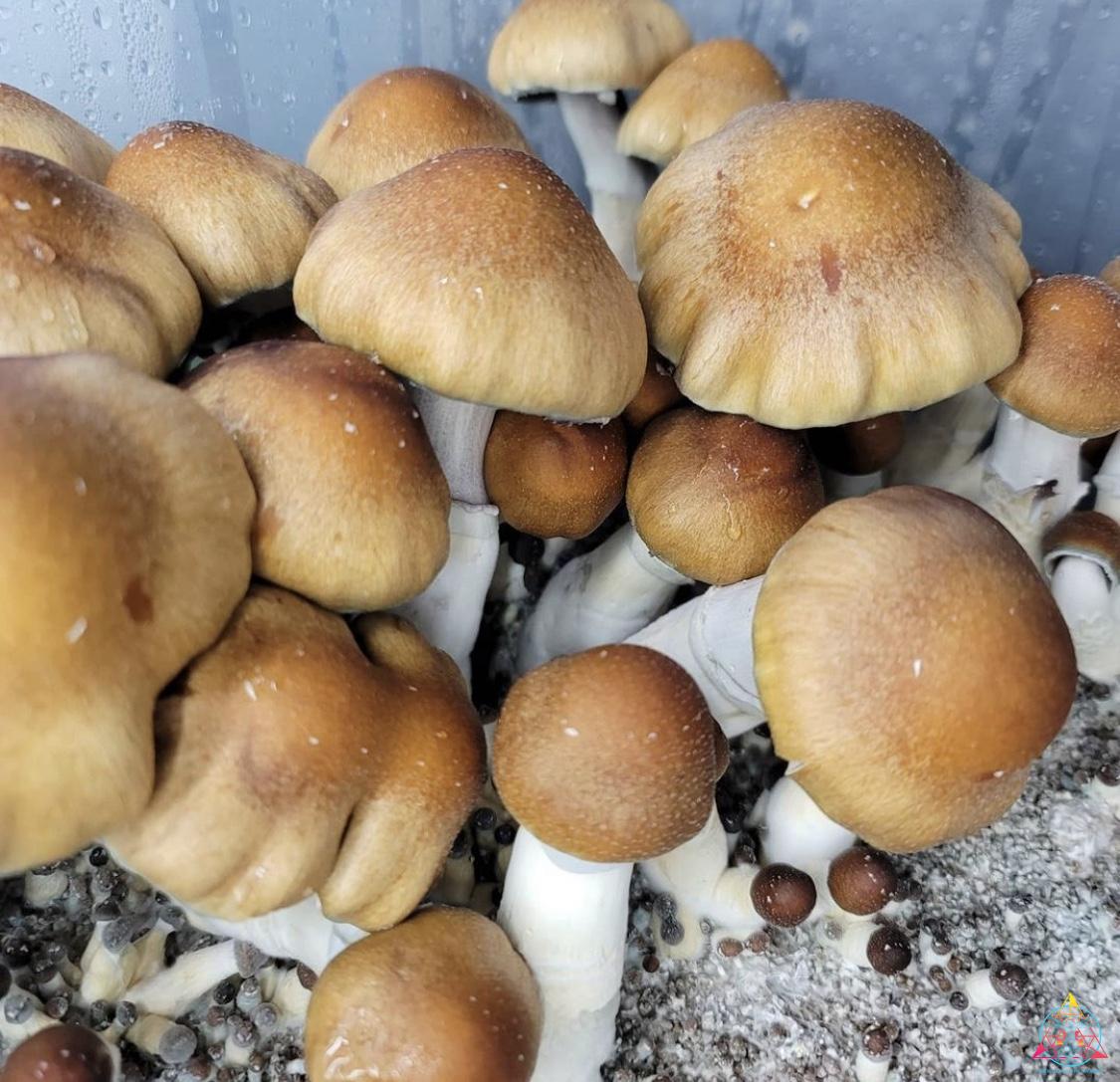
InoculateTheWorld contributed spore samples to a recent study conducted by Medicinal Genomics that aimed to further the understanding of the genes associated with psilocybin production. By sequencing and comparing 81 spore samples from numerous psilocybin-producing mushrooms, the evolution of this compound has been illuminated, all while keeping this information securely in the public domain1. Medicinal Genomics, founded in 2011 by Kevin McKernan, aimed to keep cannabis public by preventing patents on plant DNA. They also provide genetic-based testing to enhance the quality of cannabis, such as a full suite of pathogen detection tests, variant/trait screening, genetic stabilization, genetic verification, and genome sequencing. This psilocybin research is foundational for initiating similar services for the future of the mycological community. Earlier this year McKernan published a highly contiguous reference genome of the Psilocybe cubensis Penis Envy strain2. This was significant since a complete genome of P. cubensis was absent in the public NCBI database. Publishing this information prevents companies from filing for patents on that genetic material and allows access to a reference genome for future research. While this work contributes to the field, comprehension of myriad aspects about the mechanisms and evolution of the psychoactive compounds in fungi will be vital to further the cultivation industry.
Psilocybin is a secondary metabolite present in a number of phylogenetically disjointed families of Agaricales fungi (present in Bolbitiaceae, Inocybaceae, Hymenogastraceae, Pluteaceae)3. Psilocybin is a modified tryptamine derived from a common amino acid, tryptophan. After oral ingestion, psilocybin becomes rapidly dephosphorylated (removal of phosphate group by hydrolysis) to produce the compound psilocin4. Psilocin is similar in structure to Serotonin, and has an affinity for the same receptor, called the 5HT2A receptor in humans. After crossing the blood-brain barrier the compound causes notorious psychotropic effects such as altered perception, change in mood, and hallucinations5. Although psilocin and psilocybin have been classified as Schedule I substances in the United States, the narrative is beginning to change in tandem with a list of clinical applications exploring addiction treatment, depression, and end-of-life anxiety. We are still in the beginning stages of research, with limited understanding of psilocybin and its obscure analogs Baeocystin and Norbaeocystin6. Science has yet to fully grasp the mechanisms of psychoactive compounds and how their presence evolved amidst many fungi families.
The specific biosynthetic process of psilocybin production wasn’t clarified until 2017. In this research, Fricke and colleagues were able to characterize four enzymes that are necessary to transform tryptophan into psilocybin4. Biosynthesis is a multi-step, enzyme-catalyzed process where simple substrates are converted into more complex products in living organisms. Enzymes are proteins that help speed up chemical reactions in the body, and all proteins have a genetic basis since they are encoded by DNA. In the biosynthesis of psilocybin, several enzymes were recognized and labeled: PsiD functions as a new class of fungal L-tryptophan decarboxylases; PsiK catalyzes a phosphotransfer step; PsiM is a methyltransferase; PsiH is a monooxygenase. The combined reaction PsiD PsiH PsiK PsiM would be able to synthesize psilocybin from the amino acid tryptophan. Psilocybe cubensis and Psilocybe cyanescens genomes were sequenced and analyzed in order to find the genes that encode for these enzymes4. Typically, genes that are part of a biosynthesis of a particular compound are found clustered together on contiguous loci. Genes were found clustered for this enzyme synthesis and a map of the loci were produced for P. cubensis and P. cyanescens. This has been fundamental in understanding the development of psilocybin, because it then allowed scientists to search other genomes that contain this particular gene cluster. The question persists of how this gene cluster has developed and if it was inherited by different psilocybin-producing species from a common ancestor (gene homology) or evolved independently (convergent evolution).
One explanation for gene homology of psilocybin production is that psilocybin-producing species have acquired this gene cluster from horizontal gene transfer. This is when genetic material is transferred from one organism to another non-sexually, and can happen between two different species. Generally this is more common in bacteria, unicellular organisms, and simple fungi, but evidence of this phenomena in more developmentally complex organisms has recently surfaced. The actual mechanism of transfer in more complex organisms is unexplained scientifically, but still characterized as a small piece of a genome integrating into a larger genome. Horizontal gene transfer is more successful in closely related species, but can also occur in distantly related species, especially if they share a common niche or environment. Additionally, the rate of horizontal gene transfer is higher for clustered genes, especially ones that encode for enzymatic and regulatory pathways, like psilocybin. This mechanism could help explain how such disparate species from several families all produce a complex product. Research conducted by Reynolds and colleagues supports the hypothesis that lateral gene transfer of the psilocybin gene cluster is responsible for the production of psilocybin in several species3. They sequenced genomes of three distantly related psilocybin containing mushrooms, Psilocybe cyanescens, Gymnopilus dilepis and Panaeolus cyanescens, and compared these to relatively related species that don’t produce psilocybin, Galerina marginata, Gymnopilus chrysopellus, Hypholoma sublateritium respectively. A homologous multi-gene cluster was identified in all of the psilocybin producing species, and not found in all of the non-psilocybin producing species3. To confirm that these sequences are related to the psilocybin-producing enzymes, they compared the identified gene clusters to the gene cluster elucidated by Fricke and colleagues. The sequences from Psilocybe cyanescens, Gymnopilus dilepis, and Panaeolus cyanescens shared sequence similarity of 75-95% compared to the previously described gene cluster in P. cubensis3.
A similar study by Awan and colleagues analyzed genomes from psilocybin-producing species Psilocybe cyanescens, Pluteus salicinus, Gymnopilus dilepis, and Inocybe corydalina7. They also found evidence for the known biosynthesis cluster in several species, giving some support to the horizontal gene transfer hypothesis. Interestingly, the relatedness of this gene cluster correlated to the ecological environment the species inhabited. Gymnopilus dilepis is more closely related to the Psilocybe genus at the species level, but genes from the psilocybin cluster showed to be more related to the more distant relative Pluteus salicinus. Similarly, the psilocybin gene cluster is closely related between P. cubensis and Panaeolus cyanescens, although they are more distantly related species7. The close environmental association between these species could provide a mechanism for DNA transfer via horizontal gene transfer; P. cubensis and Pan. cyanescens both metabolize dung, while G. dilepis and P. salicinus live on alive or decaying wood. Although there is support for lateral gene transfer, this research also found the psilocybin gene cluster was not established in Inocybe corydalina7. This suggests an analogous biosynthetic pathway for the production of psilocybin, meaning there is identical function of the gene with independent evolution. This common process of functionally identical genes evolving separately is called convergent evolution. Despite lacking the known biosynthesis pathway, I. corydalina still produces psilocybin which has a genetic and enzymatic basis. Further analysis searched the I. corydalina genome for an alternative psilocybin gene cluster that contains all of the enzymes that convert tryptophan to psilocybin. One candidate was found, a completely novel biosynthetic cluster7. Supplementary research is required to further characterize this novel gene cluster and analyze how pervasive it is in other psilocybin-producing mushrooms.
The research completed by McKernan from Medicinal Genomics contributes to the discussion about the evolution of psilocybin biosynthesis pathway. By using a reference genome of P. cubensis strain Penis Envy, 81 genomes from several species in the Psilocybe genus were sequenced and assembled1. The majority of the genomes were a variety of popular P. cubensis strains from various cultivators, with additional species from the Psilocybe genus represented: P. tampanensis, P. galindoi, and P. azurescens. The samples Inoculate The World provided were Tosohatchee, Puerto Rican, Orissa India, Leucistic Golden Teacher, Leucistic Ecuador, Koh Samui, and AA+. All of these samples had consistently low contiguous units, or contigs, which are the number of the chunks of DNA it takes to construct the genome. Low contigs means that the chunks of DNA were longer, making them much easier to construct and read. The order of the pieces is unknown but certain areas are easy to distinguish, such as an area that codes for enzymes of a known gene cluster. When samples are contaminated, extra DNA from bacteria or fungi is added to the data that needs to be aligned. This can represent itself with more contigs that are smaller in length. Many of the spore samples sequenced had evidence of contamination, while the psilocybe spore syringe samples ITW provided were consistently clean. By searching these Psilocybe species for the presence of the psilocybin gene cluster, the research expounded upon several inquiries about psilocybin evolution. Namely, evidence was found that supports both convergent evolution and lateral gene transfer as the mechanism of psilocybin evolution1. The 81 genomes had uniform sequence coverage and high read mapping rates. Essentially, you use the reference genome to align the contiguous chunks of DNA to and high mapping rates indicate that what you sequenced is properly aligned to the reference genome. This is expected, considering most of the samples consisted of the same species as the reference genome. A small portion of the P. cubensis genome did have a surprising amount of variation in certain areas. Although these locations only contributed less than 2% of the genome, they could be important for further study1. P. tampanensis, P. galindoi, and P. azurescens exhibited the most genetic divergence from the reference genome, reiterating that they are in fact different species. These species also had low mapping rates over the psilocybin gene cluster, indicating an absence of this gene. However, these species still produce psilocybin, and must contain psilocybin-producing enzymes, which will reside somewhere in the genome. P. tampanensis, P. galindoi, and P. azurescens genomes were analyzed for different areas that might encode for the biosynthesis enzymes PsiD, PsiH, PsiK and PsiM. They found partial clusters of a PsiM gene that were not part of the previously described gene pathway and a homologous PsiK amino acid sequence was established on several alignments1. In conclusion, P. tampanensis, P. galindoi and P. azurescens showed evidence of non-contiguous, non-clustered psilocybin pathway genes. However, substantial research and characterization is needed to determine an alternate psilocybin pathway in these species.
All of the psilocybin evolution research seems to indicate that the presence of psilocybin is due to horizontal gene transfer and convergent evolution. Support for horizontal gene transfer was seen in the presence of the previously described psilocybin biosynthesis gene cluster4. This cluster was found in Psilocybe cyanescens, Gymnopilus dilepis, Pluteus salicinus, and Panaeolus cyanescens, which gives support to lateral gene transfer3 7. However, it was absent in Inocybe corydalina, Psilocybe tampanensis, Psilocybe galindoi and Psilocybe Azurescens, indicating convergent evolution1 7. All species without the known gene cluster also showed evidence for alternate psilocybin pathways. These pathways need to be further characterized, then compared to all psilocybin producing mushrooms to determine if there is pathway redundancy. Considering that many of these species are known to also contain psilocybin analogs baeocystin and norbaeocystin, it seems likely that multiple pathways exist that have yet to be identified.
Fundamentally, McKernan’s research is important because it provides a framework for the future of cultivation and breeding efforts in P. cubensis. This study initiated a catalog of the genetic variation across P. cubensis by sequencing some of the most prominent cultivated strains. The publication of these genomes helps ensure protection against private patents, which is especially important in the early stages of the psilocybin industry. Documenting the genetic variation within P. cubensis can help researchers understand the genetic basis of certain traits, compound compositions, and pathogens. Being able to associate genes to phenotypes and chemotypes could drastically improve cultivation efforts. The work by McKernan is substantial considering that a public reference genome for P. Cubensis was lacking until this year. Naturally, more characterization is required to fully understand the evolution of psilocybin in psychedelic mushrooms.
Related Articles
Psilocybe ochraceocentrata – Hidden in Plain Sight
If you follow the world of magic mushroom microscopy, you've almost certainly encountered the spores of Natal Super Strength or Transkei. They're among the most popular specimens in the hobby, studied by enthusiasts and researchers alike. Like pretty much everyone...
Are Psychedelic Retreats Worth the Trip?
“Most men and women lead lives at the worst so painful, at the best so monotonous, poor and limited that the urge to escape, the longing to transcend themselves if only for a few moments, is and has always been one of the principal appetites of the soul.” -Aldous...
The Curious Case of the Grey-Market Mushroom Edibles
The views and opinions expressed in this piece are solely those of the author, and are not representative of Inoculate the World as a company. "Magic Mushroom" Edibles are Everywhere. What's the Deal? Six years ago, very few American psychedelic users had...










